Computes posterior mean predictions from a Gaussian process regression model given precomputed kernel matrices. The implementation follows Algorithm 2.1 in Rasmussen and Williams (2006), with an optional centring step and automatic jitter escalation to handle near-singular kernel matrices.
Examples
N1 <- 25
N2 <- 10
x_train <- seq(-pi, pi, length.out = N1)
x_test <- runif(N2, -pi, pi)
distance <- outer(c(x_train, x_test), c(x_train, x_test), "-") |> abs()
kernel <- rbf(distance, length_scale = 1, variance = 1)
Kxx <- kernel[1:N1, 1:N1]
Kxstar <- kernel[(N1+1):(N1+N2), 1:N1]
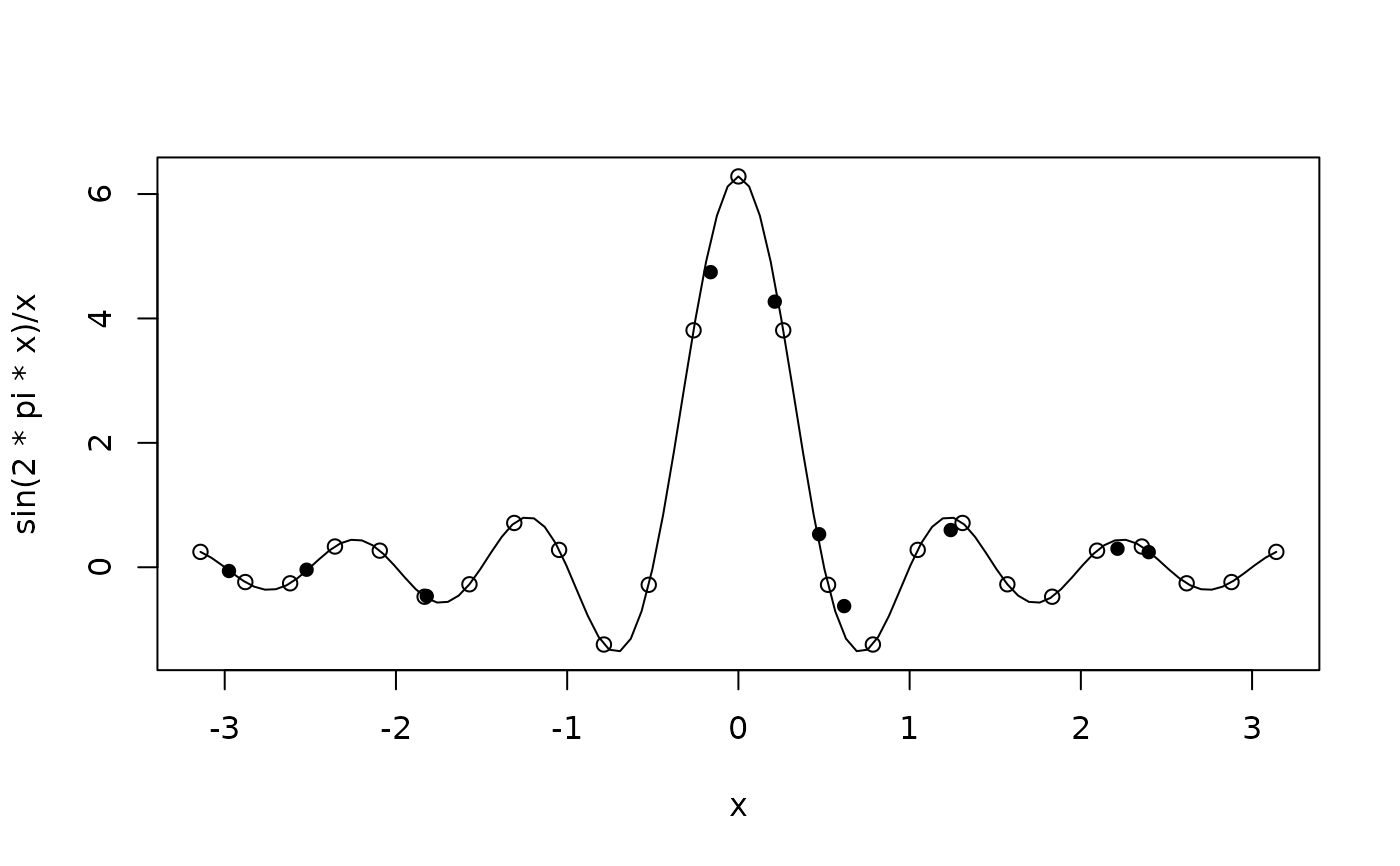
y_train <- ifelse(x_train == 0, 2*pi, sin(2*pi*x_train) / x_train)
curve(sin(2*pi*x)/x, from = -pi, to = pi)
points(x_train, y_train)
points(x_test, gpr_predict(Kxx, Kxstar, y_train), pch = 16)