(Sliced) Wasserstein - Gaussian process regression workflow
Source:vignettes/sliced-wasserstein-workflow.Rmd
sliced-wasserstein-workflow.RmdGoal
This vignette illustrates the main use of the slicer
package by means of a toy example. Empirical distributions are generated
from bivariate Gaussian mixtures with four isotropic components laid out
symmetrically around the origin. These distributions differ from one
another in that they are rotated counterclockwise by an angle
.
The goal of the analysis is to retrieve the specific angles by which the
distributions have been rotated.
Functions
Let’s load slicer.
To generate the input distributions, we define
generate_data():
generate_data <- function(n = 200, a = 5, angle = 0, Sigma = diag(1, 2)) {
mus <- cbind(c(a, 0), c(0, a), c(-a, 0), c(0, -a))
R <- rbind(
c(cos(angle), -sin(angle)),
c(sin(angle), cos(angle))
)
cluster <- sample(1:4, size = n, replace = TRUE)
d <- matrix(0, nrow = n, ncol = 2)
for (i in 1:4) {
d[which(cluster == i), ] <- MASS::mvrnorm(
sum(cluster == i), mu = mus[, i], Sigma = Sigma
)
}
d %*% t(R)
}The plots below show two empirical distributions, once using a rotation angle of and once using one of .
set.seed(2026) # for reproducibility
op <- par(no.readonly = TRUE)
par(mfrow = c(1, 2))
generate_data(angle = pi/16) |>
plot(xlab = "first dimension", ylab = "second dimension")
generate_data(angle = 3*pi/8) |>
plot(xlab = "first dimension", ylab = "second dimension")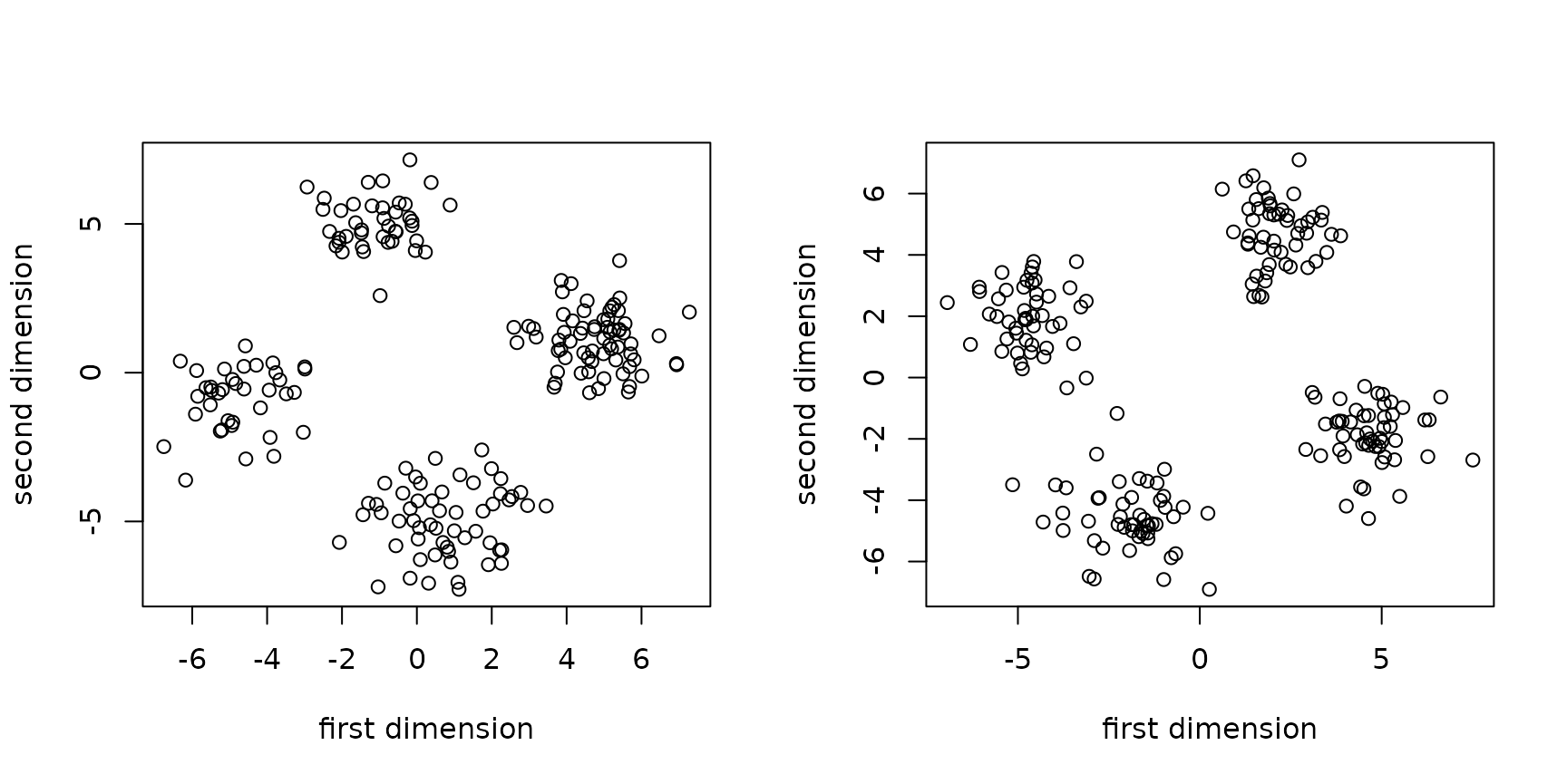
par(op)Finally, a function for computing the root mean squared error of the predictions:
Data generation
We generate 60 matrices (40 for training, 20 for testing) containing empirical distributions with the rotation angles sampled uniformly from . Each empirical distribution is based on 200 data points.
Distances
We can use compute_all_distances() to compute pairwise
distances between all 60 input distributions. (Technically, we don’t
really need the pairwise distances between the twenty inputs used for
testing.) To compute sliced Wasserstein distances, we
need to first generate
sampled uniformly at random from the unit sphere;
generate_directions() takes care of this. We’ll use
and set
since we’re working in two dimensions. If we’re only interested in the
estimated sliced Wasserstein distances, we can set
keep_projections to FALSE.
thetas <- generate_directions(L = 25, d = 2)
sw_distances <- compute_all_distances(distributions, thetas, verbose = FALSE,
keep_projections = FALSE)The
matrix sw_distances contains the squared
(estimated) sliced Wasserstein distances between the forty empirical
distributions.
We can also compute (normal) Wasserstein distances
between the distributions when they are projected along certain
dimensions. For instances, to compute the pairwise Wasserstein distances
along the first margin and along the second margin, we set the
projection directions to
and
.
We also set keep_projections = TRUE, which will cause the
function to output a list of two matrices with squared
Wasserstein distances: one for each margin.
marginal_distances <- compute_all_distances(distributions, diag(1, 2),
verbose = FALSE, keep_projections = TRUE)
str(marginal_distances)
#> List of 2
#> $ : num [1:60, 1:60] 0 1.274 0.931 0.548 0.649 ...
#> $ : num [1:60, 1:60] 0 2.177 0.121 0.445 0.599 ...Gaussian process models with tuned hyperparameters
The function fit_gpr() takes a single matrix with
squared pairwise distances and uses it as input to a Gaussian process
regression model with a Gaussian RBF kernel. The model’s and the
kernel’s hyperparameters are tuned using the training data by minimising
the negative marginal log-likelihood.
sw_fit <- fit_gpr(sw_distances,
training_idx = seq_len(N_train),
test_idx = N_train + seq_len(N_test),
y_train = angles[seq_len(N_train)],
verbose = TRUE)
#> Hyperparameter search 1 of 10.
#> Hyperparameter search 2 of 10.
#> Hyperparameter search 3 of 10.
#> Hyperparameter search 4 of 10.
#> Hyperparameter search 5 of 10.
#> Hyperparameter search 6 of 10.
#> Hyperparameter search 7 of 10.
#> Hyperparameter search 8 of 10.
#> Current optimum improved from -58.8175077 to -58.8176617.
#> Hyperparameter search 9 of 10.
#> Hyperparameter search 10 of 10.The output consists of predictions for the test data, the root mean squared error of these predictions (if the true test outcomes were provided), and the estimated hyperparameters.
str(sw_fit)
#> List of 5
#> $ test_predictions: num [1:20] 0.241 0.7091 0.2402 0.5074 0.0604 ...
#> $ RMSE : logi NA
#> $ length_scale : Named num 1.57
#> ..- attr(*, "names")= chr "length_scale"
#> $ variance : Named num 0.0758
#> ..- attr(*, "names")= chr "variance"
#> $ lambda2 : Named num 1.57e-07
#> ..- attr(*, "names")= chr "lambda2"
plot(angles[N_train + seq_len(N_test)], sw_fit$test_predictions,
xlab = "true test outcomes", ylab = "predicted test outcomes")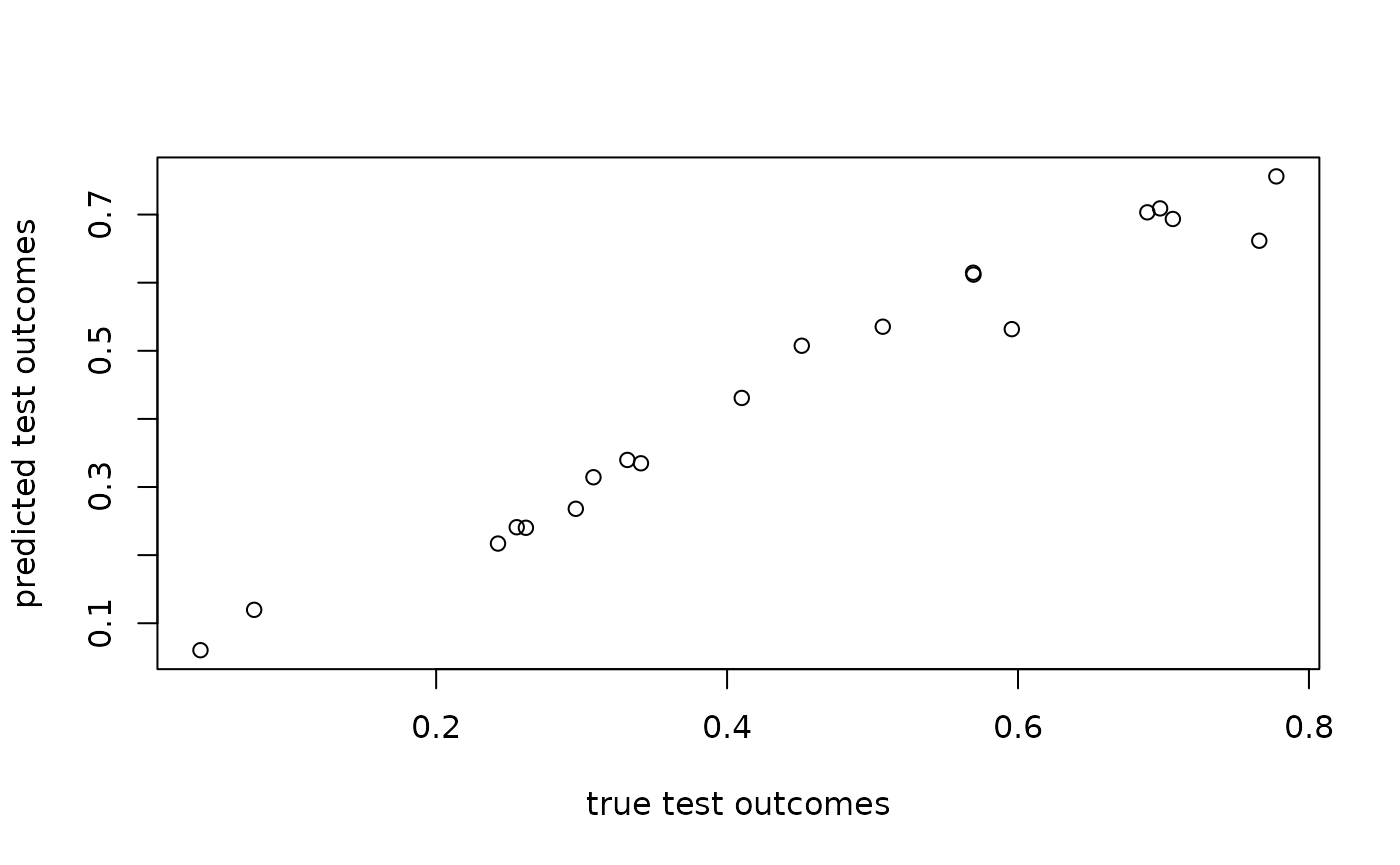
rmse(sw_fit$test_predictions, angles[N_train + seq_len(N_test)])
#> [1] 0.03804247When multiple matrices with squared distances are provided, the
function fit_gpr_multiple() can be used. Now, estimated
length-scale and (kernel) variance hyperparameters are provided for the
Gaussian RBF corresponding to each squared distance matrix.
marginal_fit <- fit_gpr_multiple(marginal_distances,
training_idx = seq_len(N_train),
test_idx = N_train + seq_len(N_test),
y_train = angles[seq_len(N_train)],
verbose = TRUE)
#> Hyperparameter search 1 of 10.
#> Hyperparameter search 2 of 10.
#> Current optimum improved from -48.3151392 to -61.5134707.
#> Hyperparameter search 3 of 10.
#> Hyperparameter search 4 of 10.
#> Hyperparameter search 5 of 10.
#> Hyperparameter search 6 of 10.
#> Hyperparameter search 7 of 10.
#> Hyperparameter search 8 of 10.
#> Current optimum improved from -61.5134707 to -61.5134707.
#> Hyperparameter search 9 of 10.
#> Hyperparameter search 10 of 10.
str(marginal_fit)
#> List of 5
#> $ test_predictions: num [1:20] 0.2395 0.6678 0.2429 0.5619 0.0919 ...
#> $ RMSE : logi NA
#> $ length_scale : Named num [1:2] 3.9 6.09
#> ..- attr(*, "names")= chr [1:2] "length_scale1" "length_scale2"
#> $ variance : Named num [1:2] 0.347 0.146
#> ..- attr(*, "names")= chr [1:2] "variance1" "variance2"
#> $ lambda2 : Named num 0.000448
#> ..- attr(*, "names")= chr "lambda2"
plot(angles[N_train + seq_len(N_test)], marginal_fit$test_predictions,
xlab = "true test outcomes", ylab = "predicted test outcomes")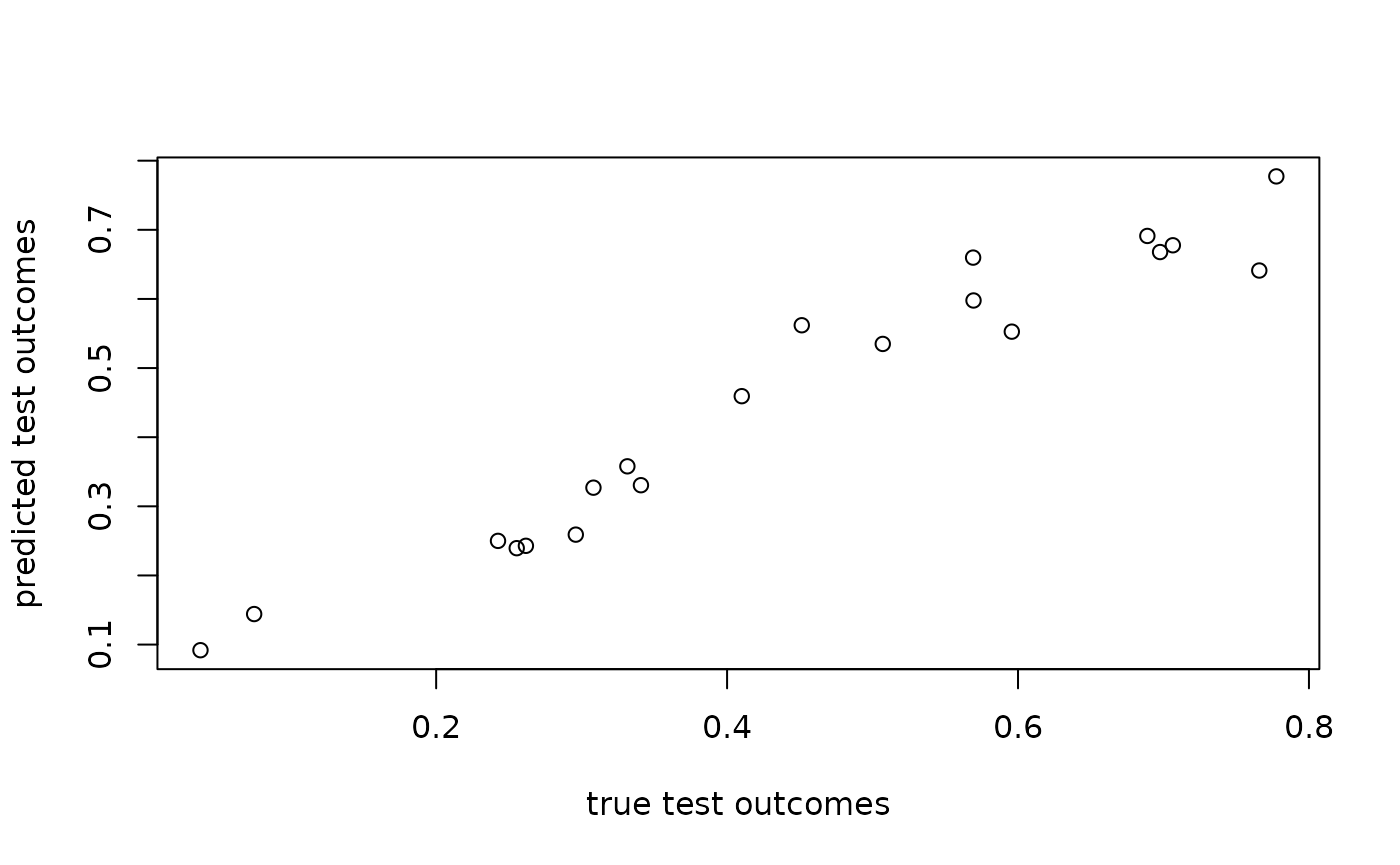
rmse(marginal_fit$test_predictions, angles[N_train + seq_len(N_test)])
#> [1] 0.05219439We can combine the marginal and sliced Wasserstein distances into a list with three distance matrices, too:
total_fit <- fit_gpr_multiple(list(sw_distances, marginal_distances[[1]], marginal_distances[[2]]),
training_idx = seq_len(N_train),
test_idx = N_train + seq_len(N_test),
y_train = angles[seq_len(N_train)],
verbose = TRUE)
#> Hyperparameter search 1 of 10.
#> Hyperparameter search 2 of 10.
#> Current optimum improved from -58.8320454 to -61.5129334.
#> Hyperparameter search 3 of 10.
#> Current optimum improved from -61.5129334 to -63.0802534.
#> Hyperparameter search 4 of 10.
#> Hyperparameter search 5 of 10.
#> Hyperparameter search 6 of 10.
#> Hyperparameter search 7 of 10.
#> Hyperparameter search 8 of 10.
#> Hyperparameter search 9 of 10.
#> Hyperparameter search 10 of 10.
str(total_fit)
#> List of 5
#> $ test_predictions: num [1:20] 0.247 0.69 0.247 0.549 0.085 ...
#> $ RMSE : logi NA
#> $ length_scale : Named num [1:3] 0.421 3.836 5.488
#> ..- attr(*, "names")= chr [1:3] "length_scale1" "length_scale2" "length_scale3"
#> $ variance : Named num [1:3] 0.00117 0.24746 0.12484
#> ..- attr(*, "names")= chr [1:3] "variance1" "variance2" "variance3"
#> $ lambda2 : Named num 3.32e-08
#> ..- attr(*, "names")= chr "lambda2"
plot(angles[N_train + seq_len(N_test)], total_fit$test_predictions,
xlab = "true test outcomes", ylab = "predicted test outcomes")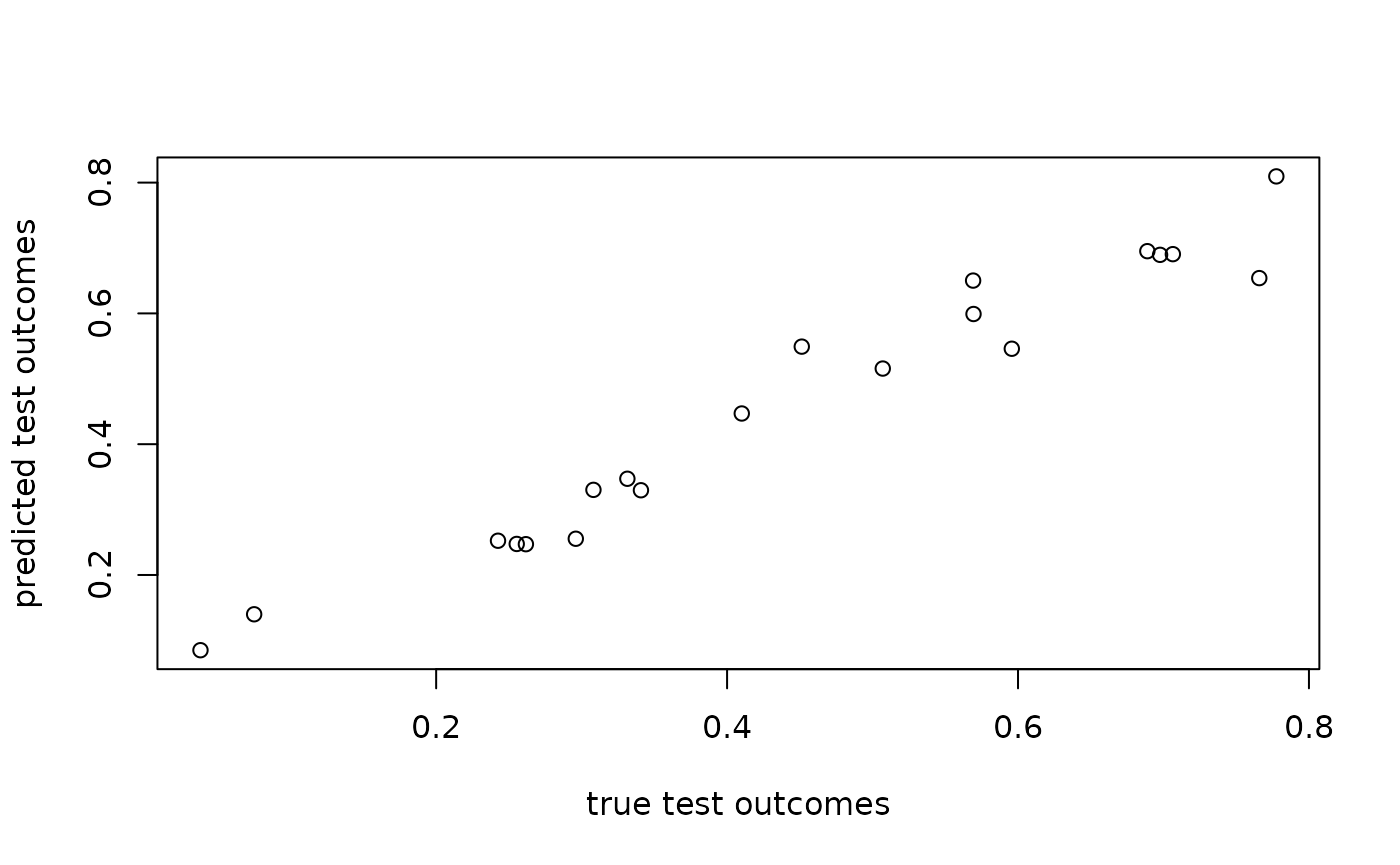
rmse(total_fit$test_predictions, angles[N_train + seq_len(N_test)])
#> [1] 0.0469791